
The Multispecific Design Toolchain is Broken
Picture this…
A scientist sits down with Excel, a spreadsheet open to a bispecific antibody template. They’ve got variable regions from a promising immunization campaign - sequences they want to stitch into an existing format design using a proven pairing technology. So they copy the first variable heavy region, paste it into the heavy chain slot, then the second into its pair. Light chains next. Four copy commands and four paste commands - eight places to silently corrupt the design. No validation caught it. No visual feedback warned them. They just created a malformed sequence that won’t work. And if they want to try ten variations? They’re repeating this manual process ten times over, each one a potential point of failure.
Current State of the MsAb Toolchain
This is how most teams are still designing multispecific antibodies (MsAbs). Not because they want to, but because there’s no tool built specifically for this workflow. Instead, they’re cobbling together an electronic lab notebook like Benchling, Dotmatics ELN, or Genedata Biologics, Excel spreadsheets, maybe PowerPoint decks or file systems full of PNG images trying to document what the format actually looks like. It’s scattered across different tools, different formats, different contexts. Nobody has a single source of truth for the design itself. But this ad hoc approach starts to completely break down the moment you move beyond a simple bispecific.
Try designing a trispecific - now you’re dealing with chain pairing complexity. Multiple distinct Fabs in a single molecule create problems, so teams often turn to single chain constructs like ScFvs or ScFabs to avoid those issues. But now you’re adding even more sequences to stitch together - heavy and light chain variable regions, linkers connecting them, then all of that getting stitched onto the rest of your format using more copy-paste in Excel. The template doesn’t scale. Your lab notebook entries don’t capture the actual structure of what you’re building. And critically, there’s no way to validate that what you’ve designed actually makes sense as a coherent structure. You’re just hoping all those sequences fit together the way you think they do. Teams hit this wall and often just stop exploring. They stick with formats they know work, because the friction of trying something new is too high.
Cost of Fragmented Workflows
The real cost of this fragmented workflow isn’t just wasted time on manual sequence stitching. It’s that teams can’t explore the design space effectively. Every new format variation, every new combination of domains or linkers or cytokines, feels like a research project in itself. So most teams converge on a handful of formats they know and stick with them. They’re not discovering better designs… they’re constrained by the tools they have. And because there’s no unified system tracking these designs, institutional knowledge gets siloed. One team’s learnings about what works don’t easily transfer to the next project. You’re constantly reinventing the wheel.
The Old Way Versus BioGlyph
In the traditional workflow, you’re moving between tools constantly. You pull sequences from your lab notebook or file system. You open Excel and manually stitch them into a template, chain by chain. You create a PowerPoint slide or PNG with a cartoon diagram of what you think the format looks like. You hope it’s right. If you want to try a variation, you start over.
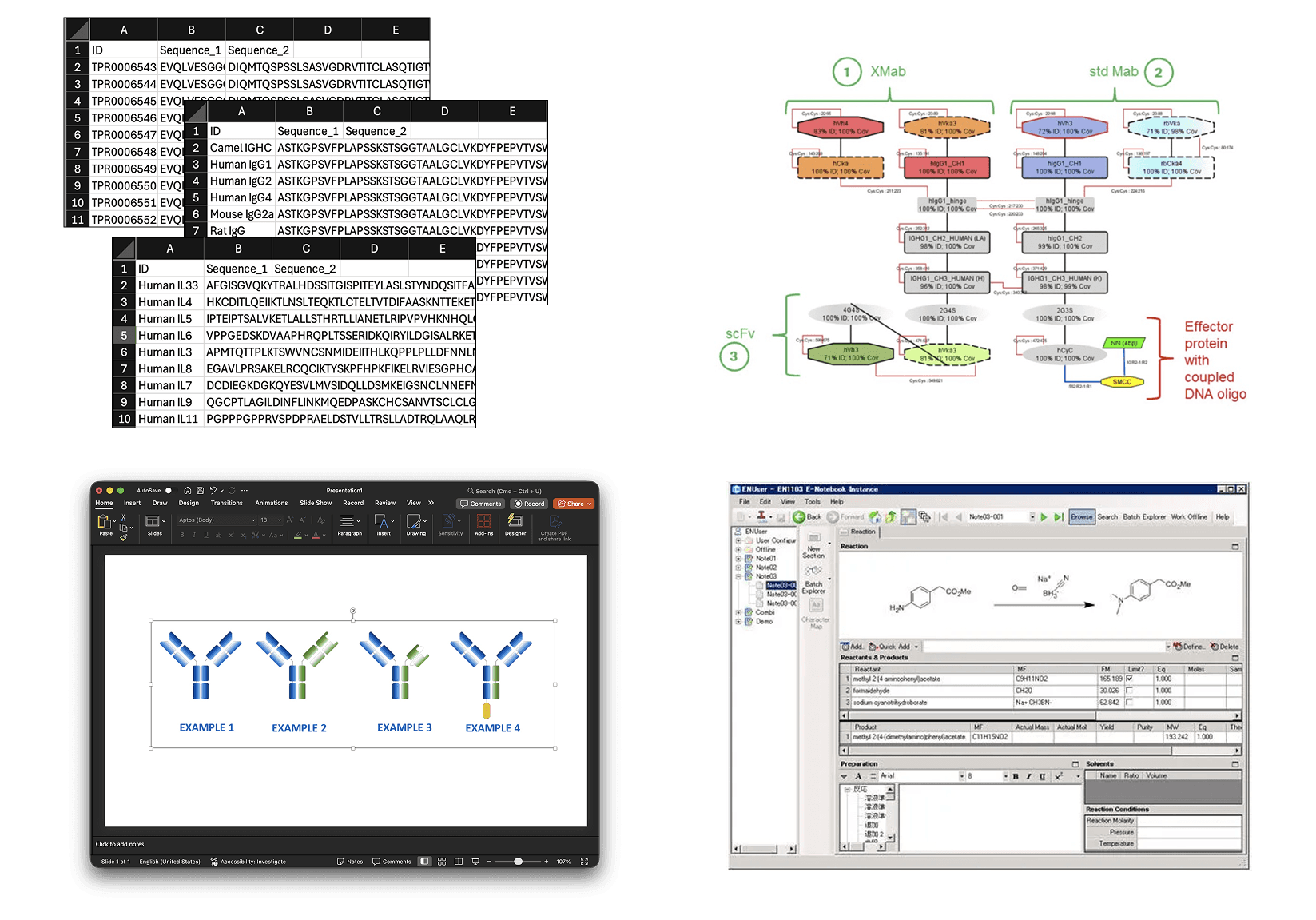

With BioGlyph, design happens in three clear stages:
Design format topologies on the design pad
Sketch your format topology on the design pad by arranging Fabs, Fcs, cytokines, linkers however you want to explore them.
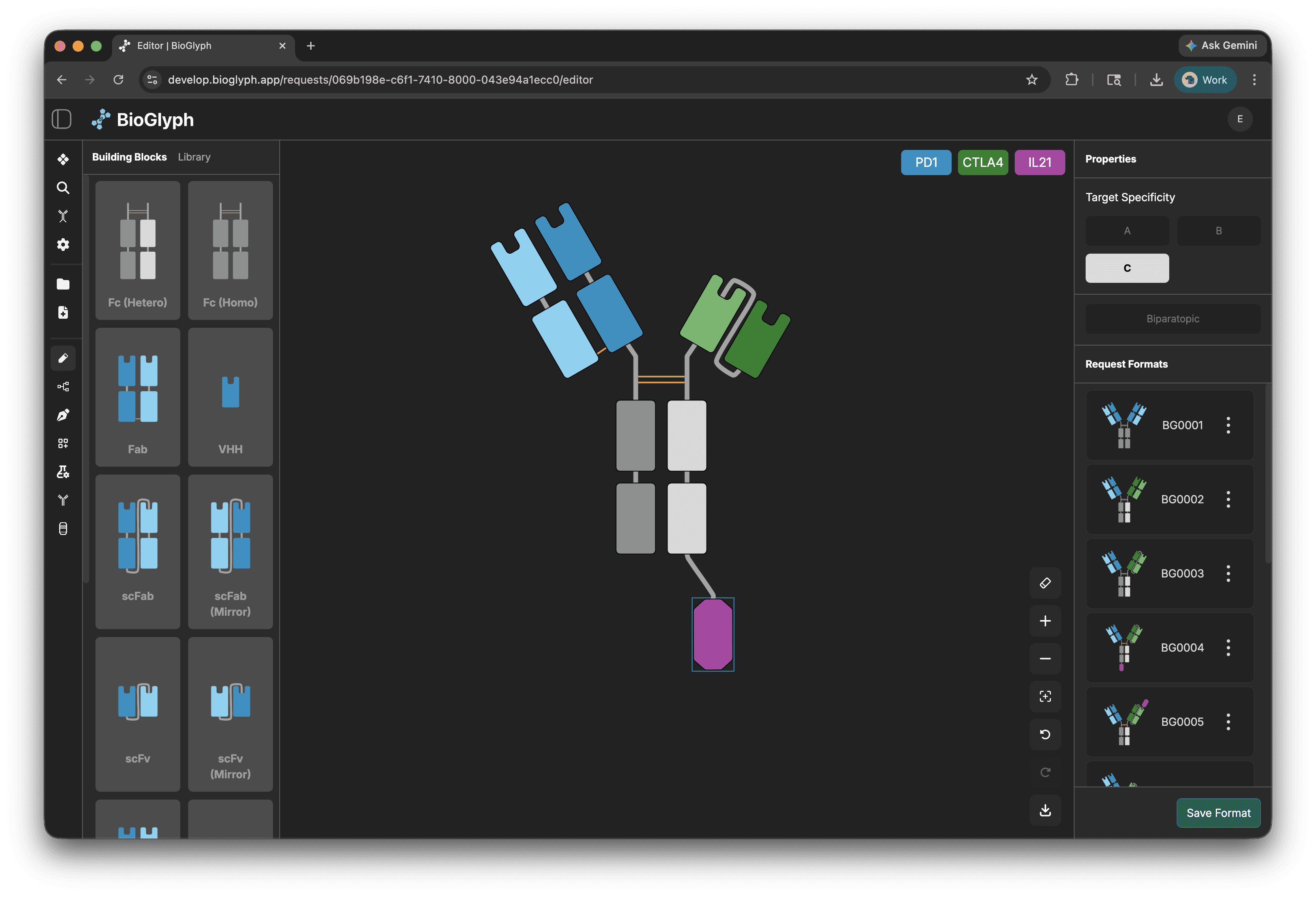
Select VRs and components in the BB panel form
In the building block panel generation form, you select your variable regions, constant regions, and linkers. You can filter variable region by property, select multiple at once, and combine with custom constant regions to create various building block types.
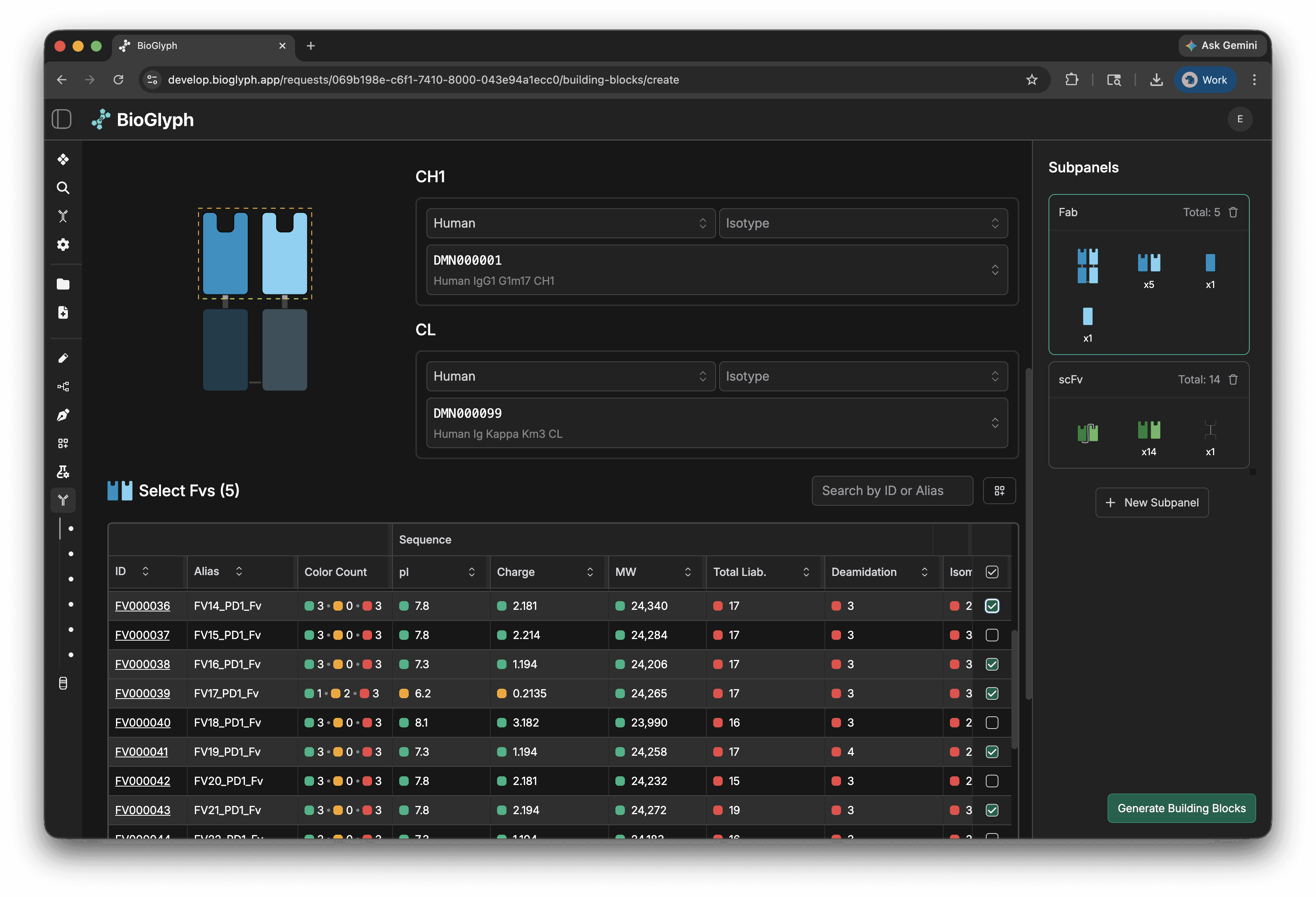
Combine BBs in the MsAb panel form
Finally, you stitch those building blocks together into your topology with optional linkers between building blocks, and the system generates all the permutations automatically.
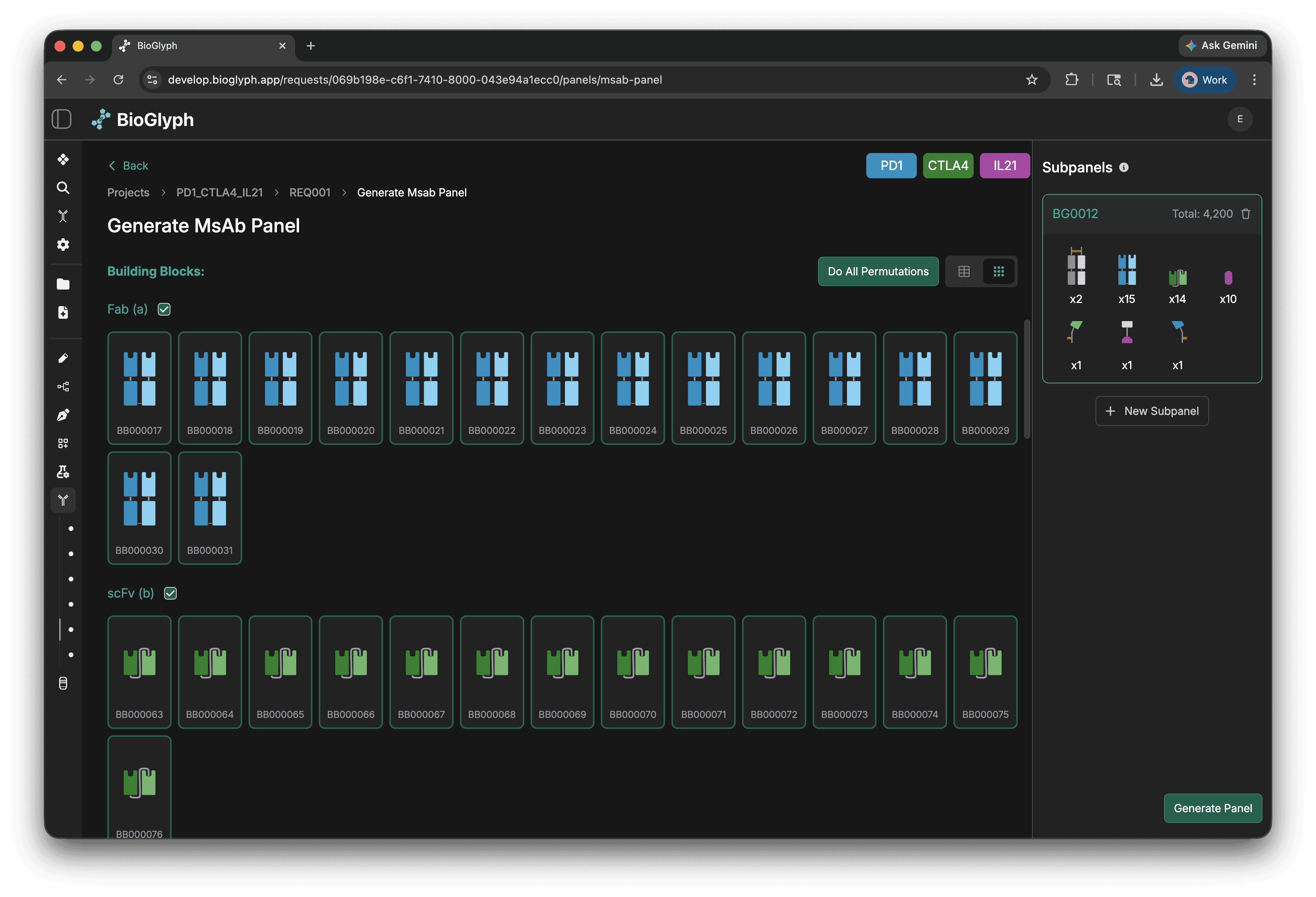
Suddenly you’re not designing one molecule at a time, you’re exploring an entire design space. One way is scattered and error-prone. The other is structured, visual, and lets you explore at scale.
A new workflow
The result is faster iteration, fewer errors, and most importantly, the ability to actually explore the multispecific design space instead of being confined to formats you know work. Teams can test hypotheses about what combinations of binding domains, linkers, and formats might yield better molecules. They’re not bottlenecked by tooling anymore. They’re limited only by their creativity and their lab capacity. That’s what a unified design platform makes possible for multispecific antibody teams.
